Welcome to Guo Lab
The PI, Xingyi Guo, PhD, is a tenured Associate Professor of Medicine in the Division of Epidemiology within the Department of Medicine at Vanderbilt University Medical Center (VUMC). Our research is inherently interdisciplinary, encompassing bioinformatics, biostatistics, AI, genetic and molecular epidemiology, population science, multi-omics integration, computational epigenetics and biology, and the use of electronic health records (EHRs) to support translational cancer research.
News & Updates editable
Paper accepted
Mixed-model and transcriptome-wide association analyses identify transcription factors and genes associated with colorectal cancer susceptibility. Nature Communications
Tool released
GLMM to detect risk TFs • sTF-TWAS • transTF-TWAS.
We’re recruiting
Openings for trainees and collaborators (see “Join”).
Research
We aim to advance the understanding of cancer etiology, prevention, and precision medicine through the development and application of bioinformatics, statistical, and machine-learning/deep-learning approaches. We integrate large-scale GWAS, multi-omics data, including single-cell and spatial omics, as well as electronic health records (EHRs) to investigate the genetic and molecular basis of human cancers. Our work focuses on identifying genetic susceptibility factors, therapeutic targets, and candidate drugs, with an emphasis on inflammatory bowel disease, colorectal adenoma, and colorectal cancer to enable precise prevention and intervention across disease progression. Dr. Guo serves as Principal Investigator or Contact PI on multiple NCI-funded studies (e.g., R37CA227130 [MERIT], R01CA269589, and R01CA297582), aimed at advancing the understanding of colorectal cancer and adenoma etiology and supporting the development of therapeutic strategies for disease prevention and intervention.
1) Genomics & disease risk
Identify susceptibility genes, fine-map loci, and build interpretable risk models across diverse populations.
- GWAS / PRS / rare variant analyses
- Functional annotation and prioritization
- Cross-ancestry evaluation
2) Multi-omics integration
Integrate transcriptomics, epigenomics, proteomics, and clinical phenotypes to identify pathways and therapeutic hypotheses.
- TWAS / PWAS / MeWAS
- Network and pathway modeling
- Colocalization / mediation analyses
3) Colorectal adenoma cohort building
Develop EHR-linked phenotypes and conduct biobank-based genomic studies of colorectal adenoma and recurrence (PIs:Guo/Yin, R01CA297582).
- Clinical cohort building and phenotyping
- Demographic and clinicopathologic risk factors
- Large-scale GWAS / WGS for adenoma
4) EHR, drug repurposing & functional validation
Integrate GWAS, proteomics, and EHRs to identify druggable proteins and therapeutic candidates for cancer prevention.
- Drug–protein data integration
- Trial emulation and real-world evidence
- Experimental validation of candidates
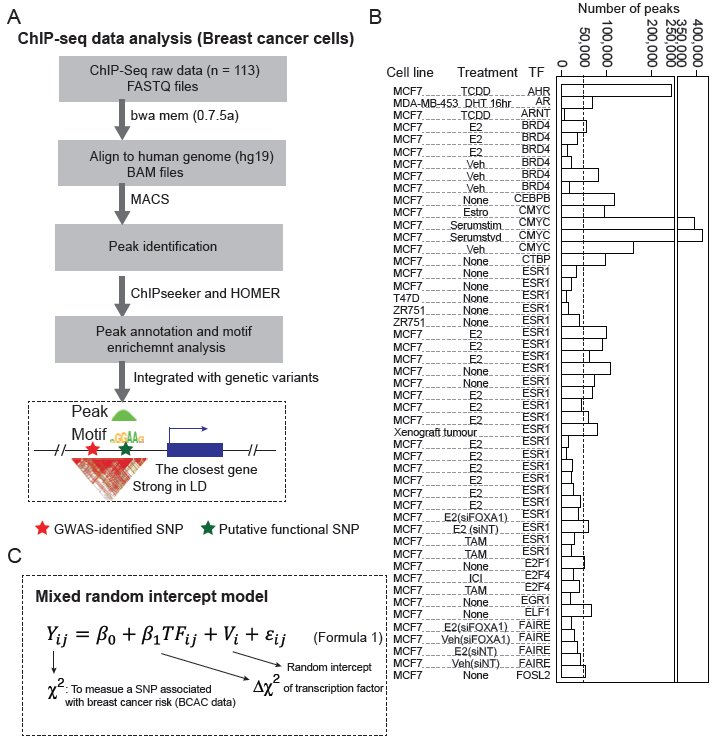
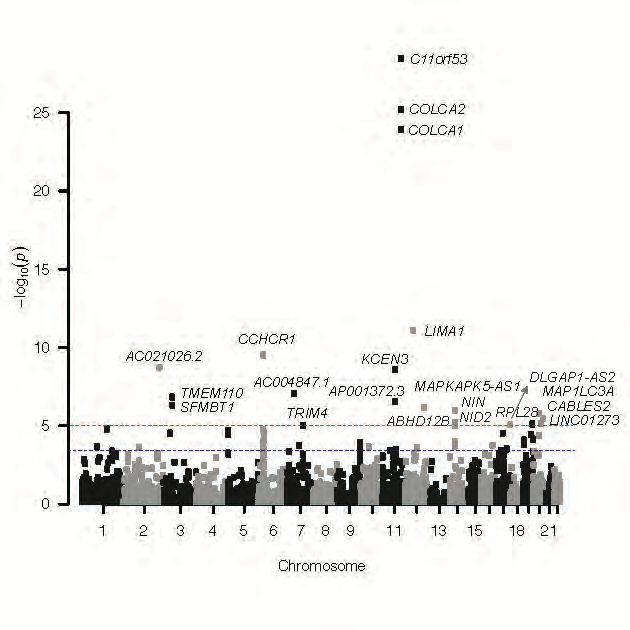
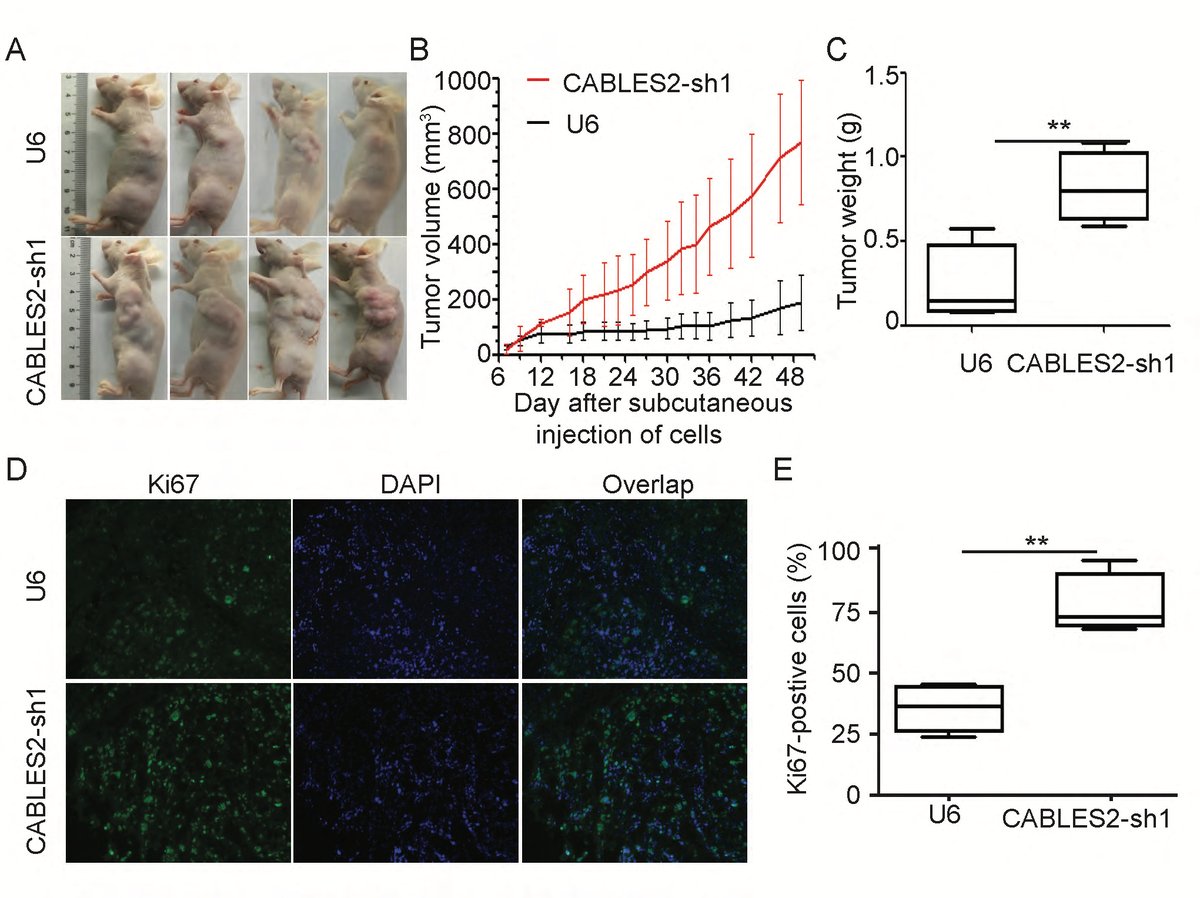
Research Highlights
Selected figures illustrating recent projects and methods.





Tools & Resources
We build and maintain software for statistical analysis to improve discovery of risk genes and transcription factors.
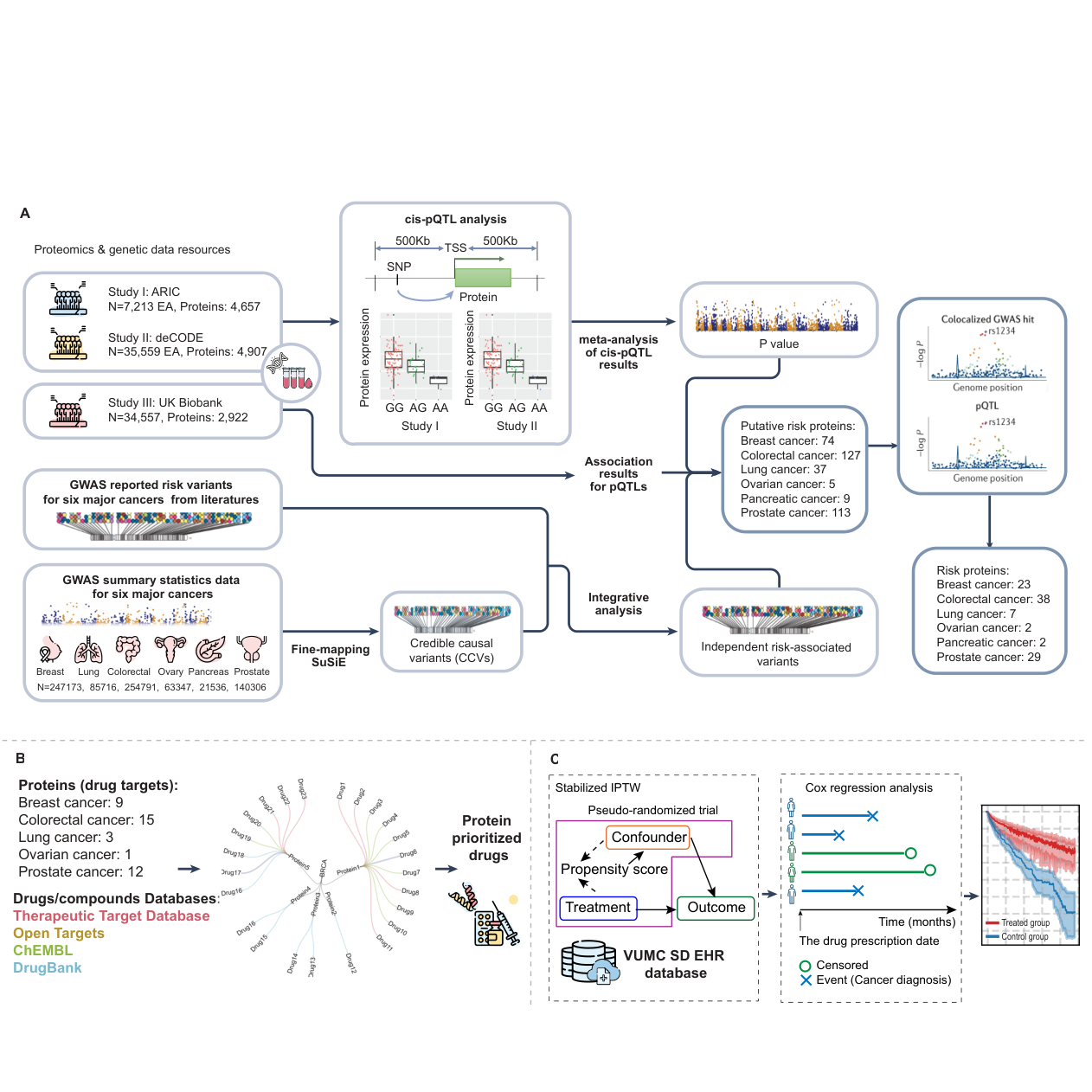
PQTL_EHR framework Workflow
Integrates cancer GWAS, proteomics, and EHRs to identify druggable proteins and therapeutic candidates.
Colorectal adenoma GWAS Dataset
Resources and summary results (add public portal link if available).
Team
Current members.
Publications
Selected publications from the Guo Lab (* corresponding author).
-
Large-scale integration of omics and electronic health records to identify potential risk protein biomarkers and therapeutic drugs for cancer prevention.
-
Demographic and Clinicopathologic Factors Associated with Colorectal Adenoma Recurrence.
-
Mixed-model and transcriptome-wide association analyses identify transcription factors and genes associated with colorectal cancer susceptibility.
-
Enhancing disease risk gene discovery by integrating transcription factor-linked trans-variants into transcriptome-wide association analyses.
-
Large-Scale Alternative Polyadenylation-Wide Association Studies to Identify Putative Cancer Susceptibility Genes.
-
Novel insights into genetic susceptibility for colorectal cancer from transcriptome-wide association and functional investigation.
-
Racial/Ethnic and Sex Differences in Somatic Cancer Gene Mutations among Patients with Early-Onset Colorectal Cancer.
-
Integrating transcription factor occupancy with transcriptome-wide association analysis identifies susceptibility genes in human cancers.
-
Distinct Genomic Landscapes in Early-Onset and Late-Onset Endometrial Cancer.
-
Genetic variations of DNA bindings of FOXA1 and co-factors in breast cancer susceptibility.
-
Identifying Novel Susceptibility Genes for Colorectal Cancer Risk From a Transcriptome-Wide Association Study of 125,478 Subjects.
-
Spectrum of Somatic Cancer Gene Variations Among Adults With Appendiceal Cancer by Age at Disease Onset.
-
Identifying Putative Susceptibility Genes and Evaluating Their Associations with Somatic Mutations in Human Cancers.
-
Discovery of rare coding variants in OGDHL and BRCA2 in relation to breast cancer risk in Chinese women.
-
Use of deep whole-genome sequencing data to identify structure risk variants in breast cancer susceptibility genes.
-
A Comprehensive cis-eQTL Analysis Revealed Target Genes in Breast Cancer Susceptibility Loci Identified in Genome-wide Association Studies.
Join the Lab
We welcome motivated trainees and collaborators. If you’re interested, please email a CV and a short description of your interests, preferred start date, and relevant experience.
Open roles for PhD students/postdocs
- Bioinformatics, Statistics, Computer Science
- Genomics, Multi-omics, EHR methods
- Analytical Pipelines, Cancer Biology
What to include
- CV + links (e.g., GitHub)
- 1–2 paragraphs: your interests & fit
- Example projects or papers
Contact
Address
Departments of Medicine & Biomedical Informatics
Vanderbilt University Medical Center
2525 West End Ave
Nashville, TN 37203